Research
The Healthy Aging Study

The Healthy Aging Study is a collaboration between the Genome Sciences Centre and other leading research groups investigating the individual factors which contribute to healthy aging and resistance to age-related disease.
Colorectal Cancer Screening

Colorectal cancer is the second leading cause of cancer-related death in Canada. The goal of this program is to develop novel easy-to-administer tests that identify individuals at the earliest, most curable stages of the disease.
Autophagy
The study of autophagy in human health is a research field that has generated tremendous attention due to the recognition that autophagy is involved in multiple developmental processes and various human diseases including cancer.
Antimicrobial Peptide Discovery
A collaborative project in Dr. Birol’s group is focusing on short proteins called antimicrobial peptides (AMPs), which are produced naturally by various animal and plant species.
Cancer Bioinformatics
We aim to use bioinformatics to investigate the landscape of mutations present in cancer genomes and the early genomic events that give rise to and promote the progression of cancer.
Cancer Epigenetics
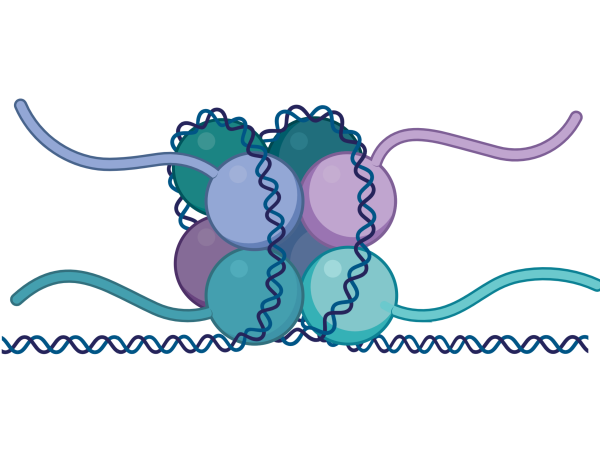
We are working to understand the role of epigenetics in cancer and to investigate the therapeutic potential of interventions directed at epigenetic processes.
Aberrant RNA Processing in Cancer
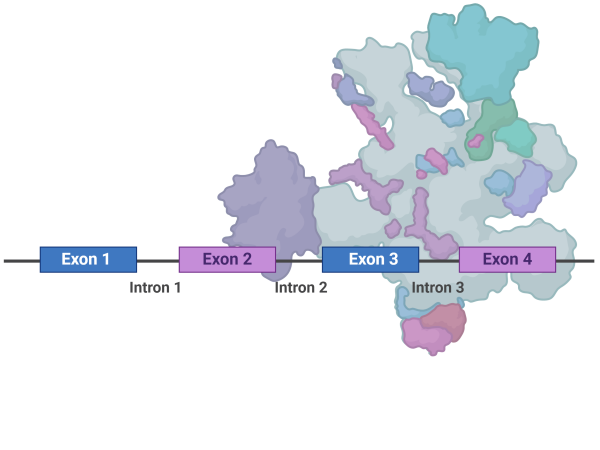
A major research focus is the regulatory mechanisms of aberrant RNA processing in cancer, particularly the regulation of alternative pre-mRNA splicing and processing, and miRNA biogenesis.
Lymphoid Cancer Family Studies
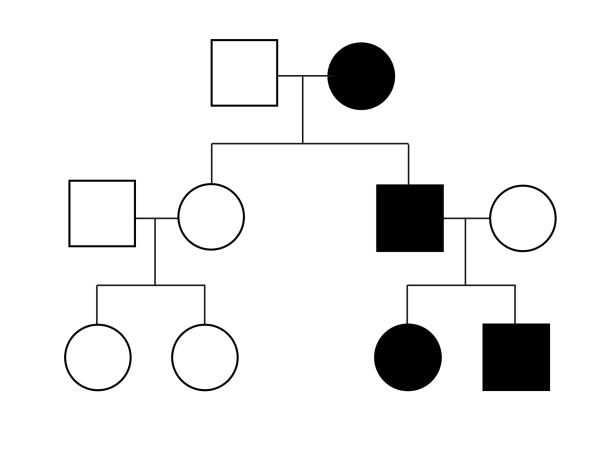
We have undertaken a family-based study to identify genetic factors contributing to lymphoid cancers including lymphoma, leukemia and myeloma.
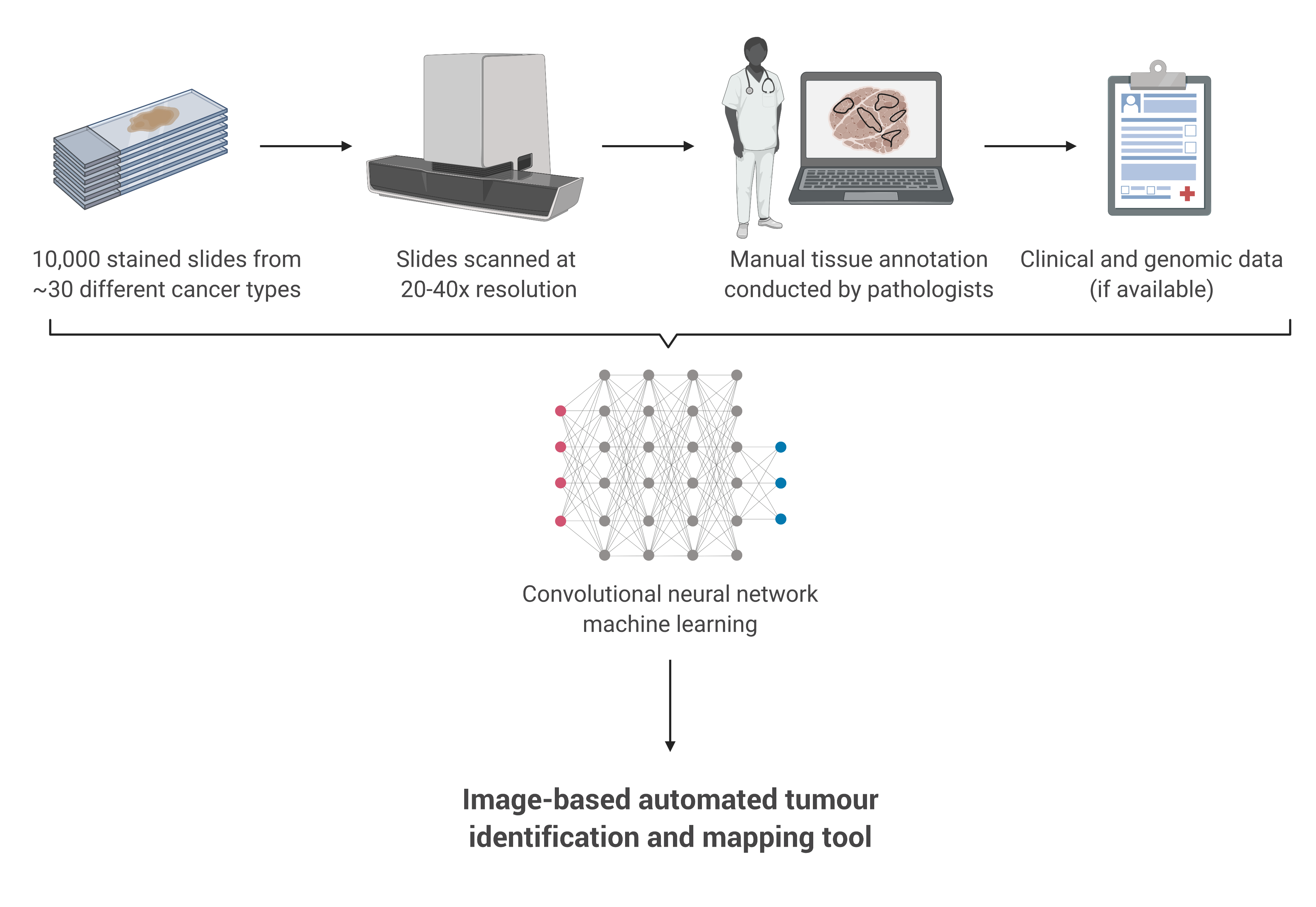
Cancer Genomes and Epigenomes
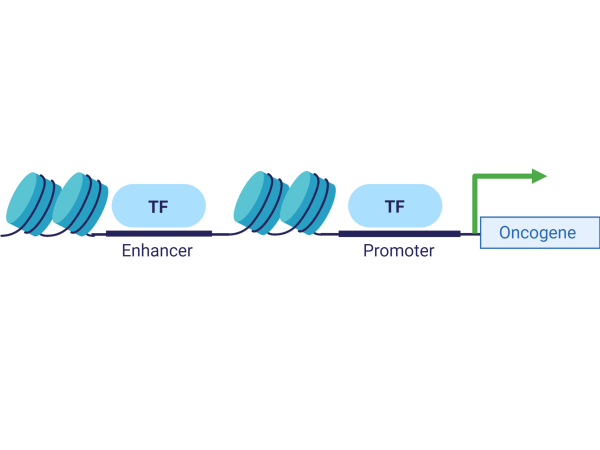
Immunotherapy
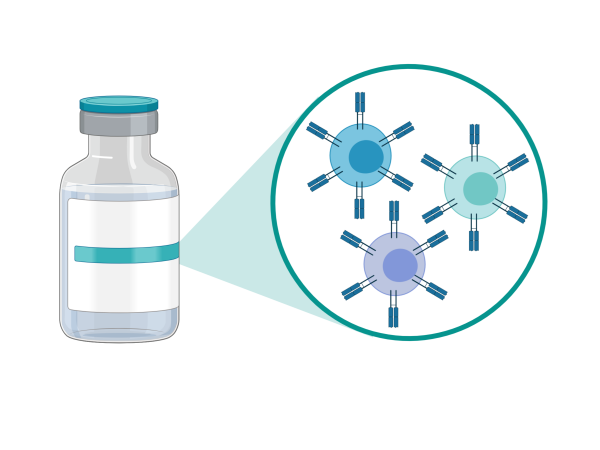
Cancer immunotherapies using engineered autologous T cells have shown remarkable efficacy against some cancer. Dr. Holt’s team is engineering T cells to selectively deliver modified cytotoxic payloads and pro-drug activators for the purpose of enhanced tumour cell killing and overcoming immune resistance.
Cancer and Infectious Agents
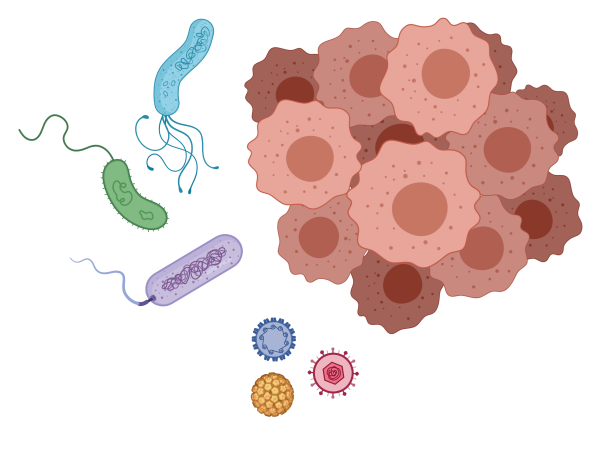
A substantial proportion (at least 15 per cent) of the global cancer burden is attributable to known infectious agents. The Holt Lab finds pathogens by their sequence signatures in human tissues using genomic methods, which revealed a strong link between colorectal carcinoma and Fusobacterium nucleatum.
Cancer Drug Resistance
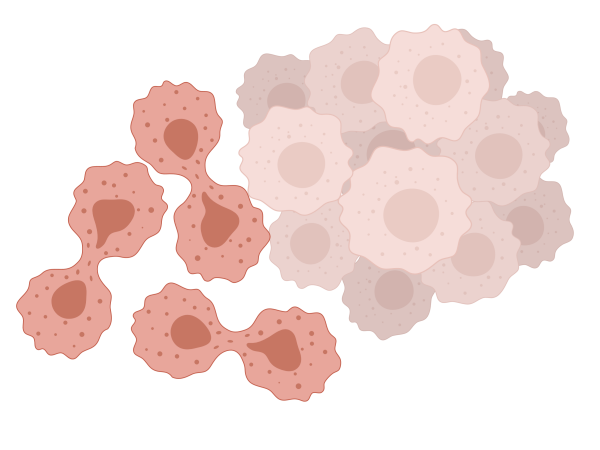
The aim of this project is to understand the mechanisms of Lenalidomide resistance in Myelodysplastic Syndromes (MDS), as well as to potentially provide strategies to overcome or bypass resistance.
Burkitt Lymphoma
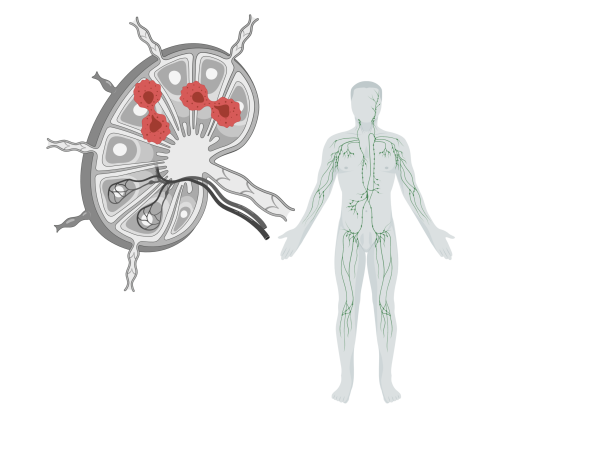
This project seeks to generate high-throughput whole genome and transcriptome sequencing data from over 100 Burkitt lymphoma tumours in order to elucidate the genetic and molecular underpinnings of the disease.
Non-coding Mutations in NHLs
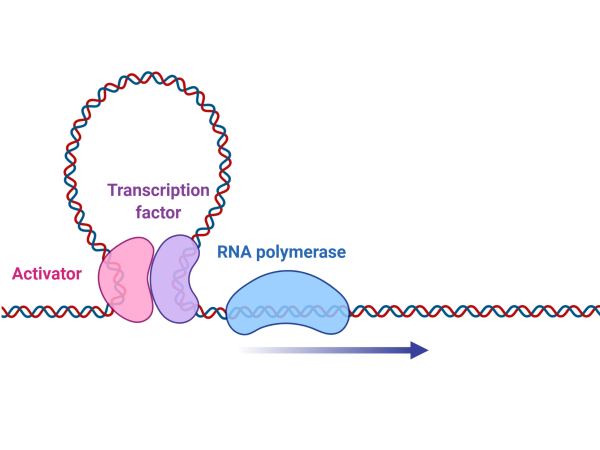
We are seeking to identify driver mutations that contribute to some features of Non Hodgkin Lymphomas using data-driven comprehensive methods that allow identification of non-coding mutations relevant to malignancy.
Novel Prostate Cancer Therapies
The Sadar lab has identified the N-terminal domain of the androgen receptor as a novel therapeutic target for the development of novel prostate cancer therapeutics.
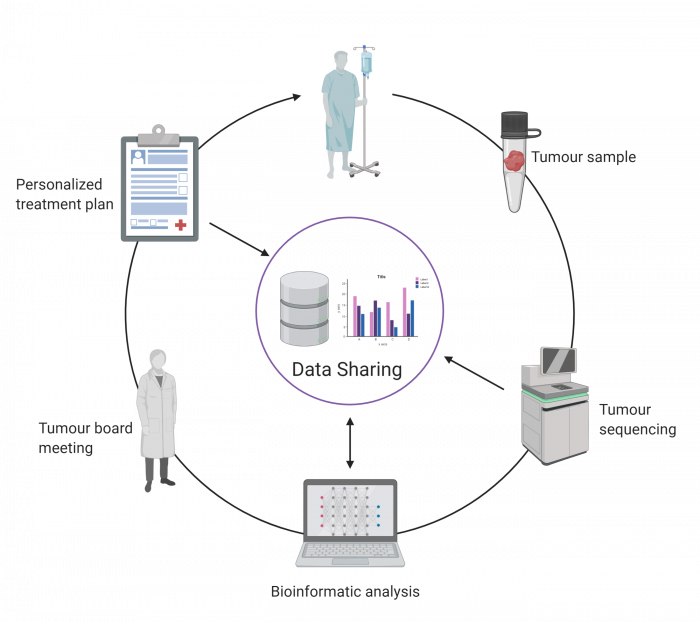 A key research objective for the POG program is to identify and better understand the genomic alterations that drive cancer and its progression and
A key research objective for the POG program is to identify and better understand the genomic alterations that drive cancer and its progression and