Distributions and lists of all significant differential expression values of the type: ActiveIntergenicRegion for comparisons: A431_IRTR_vs_DMSO, PC9_IRTR_vs_DMSO (p-value < 0.05 after multiple testing correction and Fold-Change > 1.5)
Distribution of log2 differential expression values for comparison: A431_IRTR_vs_DMSO and data type: ActiveIntergenicRegion
Distribution of all differential expression values that meet the p-value and fold-change cutoff for the feature type: ActiveIntergenicRegion. The total number of significant DE features, as well as the max and min log2 DE observed are noted in the legend. *If you can not see the figure below, click here
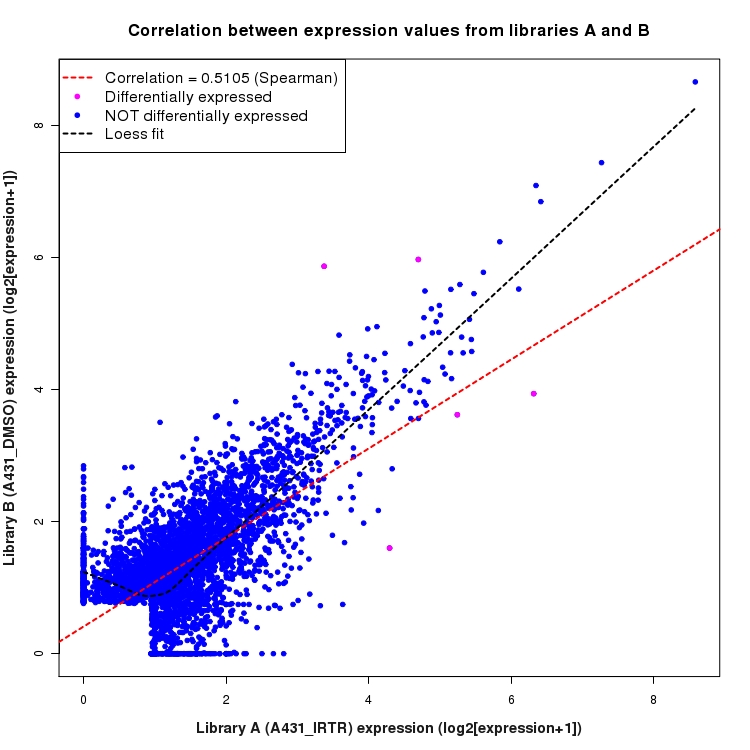
Scatter plot of log2 expression values for comparison: A431_IRTR_vs_DMSO and data type: ActiveIntergenicRegion
Correlation between expression values for all features that are expressed above background in one or both libraries for the feature type: ActiveIntergenicRegion. Features that are differentially expressed (meet the p-value and fold-change cutoff) are indicated in magenta. A linear model is fit to the data and the correlation by Spearman method is reported. A loess model is also fit to illustrate the trend of the data.
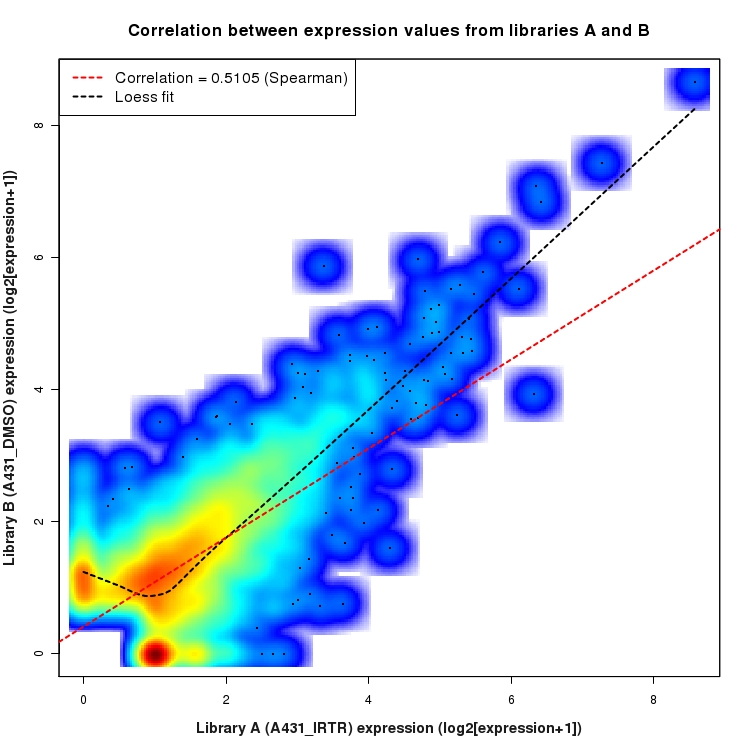
SmoothScatter plot of log2 expression values for comparison: A431_IRTR_vs_DMSO and data type: ActiveIntergenicRegion
Correlation between expression values for all features that are expressed above background in one or both libraries for the feature type: ActiveIntergenicRegion. A linear model is fit to the data and the correlation by Spearman method is reported. A loess model is also fit to illustrate the trend of the data.
Distribution of log2 differential expression values for comparison: PC9_IRTR_vs_DMSO and data type: ActiveIntergenicRegion
Distribution of all differential expression values that meet the p-value and fold-change cutoff for the feature type: ActiveIntergenicRegion. The total number of significant DE features, as well as the max and min log2 DE observed are noted in the legend. *If you can not see the figure below, click here
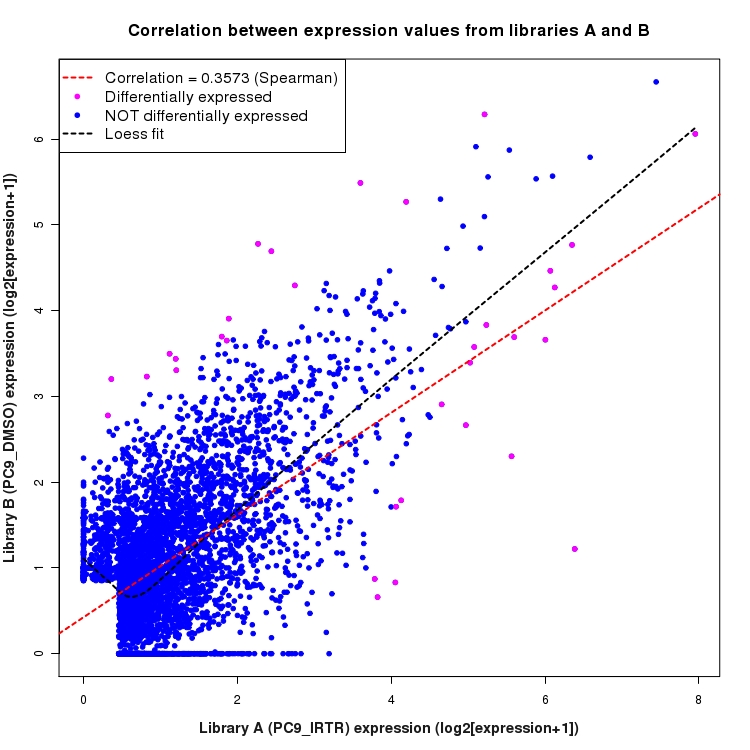
Scatter plot of log2 expression values for comparison: PC9_IRTR_vs_DMSO and data type: ActiveIntergenicRegion
Correlation between expression values for all features that are expressed above background in one or both libraries for the feature type: ActiveIntergenicRegion. Features that are differentially expressed (meet the p-value and fold-change cutoff) are indicated in magenta. A linear model is fit to the data and the correlation by Spearman method is reported. A loess model is also fit to illustrate the trend of the data.
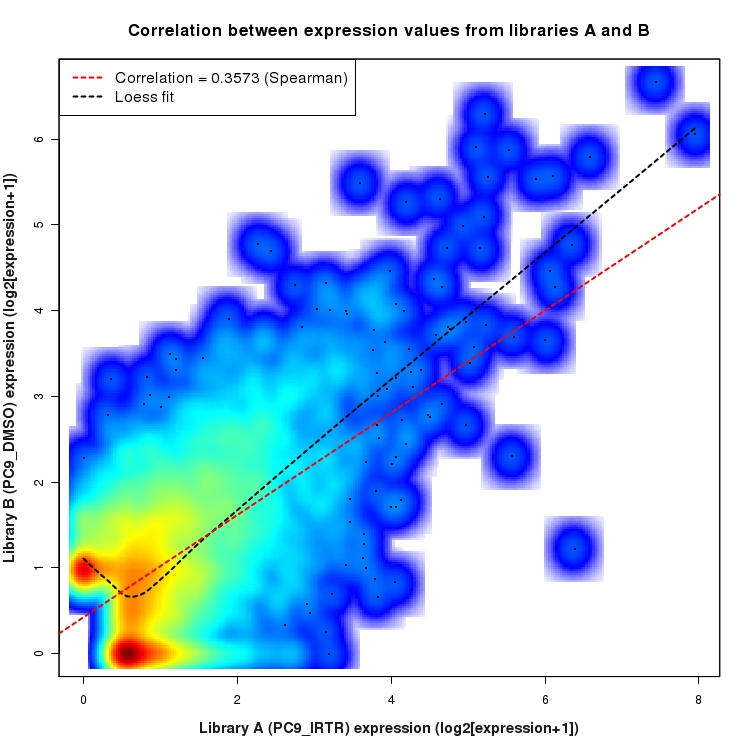
SmoothScatter plot of log2 expression values for comparison: PC9_IRTR_vs_DMSO and data type: ActiveIntergenicRegion
Correlation between expression values for all features that are expressed above background in one or both libraries for the feature type: ActiveIntergenicRegion. A linear model is fit to the data and the correlation by Spearman method is reported. A loess model is also fit to illustrate the trend of the data.
Significant differentially expressed ActiveIntergenicRegion features
The following table provides a ranked list of all significant differentially expressed ActiveIntergenicRegion features. Each column corresponds to a pair-wise comparison of two libraries. Each table cell contains the Gene Name (which links to the ALEXA-Seq gene record), the Feature Name, the name of each library being compared (which links to the feature's coordinates in the UCSC Genome Browser and displays expression data), the fold-change (FC) of the differential expression event, and the multiple testing corrected p-value for the feature. A bold row indicates that the feature is not currently supported by EST or mRNA sequence alignments. For exon junction features the number of exons skipped by the junction is indicated as 'Sn' where n is the number of exons skipped (e.g. S0 means no exons skipped, S1 means one exon skipped, etc.).
| RANK | A431_IRTR_vs_DMSO (Gene | Feature | Links | Values) | PC9_IRTR_vs_DMSO (Gene | Feature | Links | Values) |
| 1 | TRIM31 | IG51_AR1 | (A431_IRTR) | (A431_DMSO) | FC = 6.46 | q.value = 0.0175 | DHRS3 AADACL4 | IG45_AR1 | (PC9_IRTR) | (PC9_DMSO) | FC = 35.85 | q.value = 5e-26 |
| 2 | Y_RNA U6 | IG16_AR11 | (A431_IRTR) | (A431_DMSO) | FC = -5.64 | q.value = 9.87e-08 | C14orf73 TNFAIP2 | IG11_AR3 | (PC9_IRTR) | (PC9_DMSO) | FC = 9.59 | q.value = 1.42e-09 |
| 3 | DHRS3 AADACL4 | IG45_AR1 | (A431_IRTR) | (A431_DMSO) | FC = 5.18 | q.value = 1.72e-11 | Y_RNA LIMCH1 | IG42_AR1 | (PC9_IRTR) | (PC9_DMSO) | FC = 9.34 | q.value = 0.012 |
| 4 | TAF8 C6orf132 | IG21_AR11 | (A431_IRTR) | (A431_DMSO) | FC = 3.08 | q.value = 0.0273 | CALM2 TACSTD1 | IG40_AR19 | (PC9_IRTR) | (PC9_DMSO) | FC = 8.96 | q.value = 0.0366 |
| 5 | LAMC2 NMNAT2 | IG29_AR1 | (A431_IRTR) | (A431_DMSO) | FC = -2.42 | q.value = 0.0258 | PNPLA2 EFCAB4A | IG36_AR2 | (PC9_IRTR) | (PC9_DMSO) | FC = 7.54 | q.value = 0.0463 |
| 6 | EMPTY | AC091544.11 CHD2 | IG50_AR24 | (PC9_IRTR) | (PC9_DMSO) | FC = -7.17 | q.value = 0.00406 |
| 7 | EMPTY | SLC45A3 NUCKS1 | IG22_AR6 | (PC9_IRTR) | (PC9_DMSO) | FC = -5.70 | q.value = 7.13e-07 |
| 8 | EMPTY | SRD5A1 POLS | IG48_AR8 | (PC9_IRTR) | (PC9_DMSO) | FC = -5.51 | q.value = 0.0394 |
| 9 | EMPTY | AC084262.6 DUSP4 | IG28_AR2 | (PC9_IRTR) | (PC9_DMSO) | FC = -5.31 | q.value = 0.0309 |
| 10 | EMPTY | NRBP2 PLEC1 | IG44_AR4 | (PC9_IRTR) | (PC9_DMSO) | FC = -5.20 | q.value = 0.017 |
| 11 | EMPTY | DCPS ST3GAL4 | IG42_AR8 | (PC9_IRTR) | (PC9_DMSO) | FC = 5.09 | q.value = 0.0366 |
| 12 | EMPTY | ALDH3B1 NDUFS8 | IG6_AR1 | (PC9_IRTR) | (PC9_DMSO) | FC = 5.07 | q.value = 4.86e-09 |
| 13 | EMPTY | RP11-389K14.2 DIRAS2 | IG18_AR12 | (PC9_IRTR) | (PC9_DMSO) | FC = 5.07 | q.value = 0.0309 |
| 14 | EMPTY | SPAG9 | IG1_AR3 | (PC9_IRTR) | (PC9_DMSO) | FC = 4.94 | q.value = 0.000282 |
| 15 | EMPTY | SEPT11 CCNI | IG30_AR8 | (PC9_IRTR) | (PC9_DMSO) | FC = -4.76 | q.value = 7.29e-06 |
| 16 | EMPTY | RLBP1L1 ASPH | IG21_AR1 | (PC9_IRTR) | (PC9_DMSO) | FC = -4.72 | q.value = 0.017 |
| 17 | EMPTY | SMUG1 CBX5 | IG24_AR2 | (PC9_IRTR) | (PC9_DMSO) | FC = -4.29 | q.value = 0.0375 |
| 18 | EMPTY | AC010336.8 AC010336.8 | IG27_AR2 | (PC9_IRTR) | (PC9_DMSO) | FC = -4.05 | q.value = 0.012 |
| 19 | EMPTY | PDE4DIP SEC22B | IG24_AR2 | (PC9_IRTR) | (PC9_DMSO) | FC = 3.75 | q.value = 5.52e-05 |
| 20 | EMPTY | ZNF524 ZNF784 | IG3_AR5 | (PC9_IRTR) | (PC9_DMSO) | FC = -3.73 | q.value = 0.0355 |
| 21 | EMPTY | SSU_rRNA_5 | IG1_AR1 | (PC9_IRTR) | (PC9_DMSO) | FC = 3.71 | q.value = 5.86e-30 |
| 22 | EMPTY | hsa-mir-663 RP4-610C12.2 | IG23_AR3 | (PC9_IRTR) | (PC9_DMSO) | FC = -3.71 | q.value = 7.31e-08 |
| 23 | EMPTY | TRIM56 SERPINE1 | IG11_AR6 | (PC9_IRTR) | (PC9_DMSO) | FC = 3.62 | q.value = 1.96e-07 |
| 24 | EMPTY | MAP2K7 SNAPC2 | IG31_AR7 | (PC9_IRTR) | (PC9_DMSO) | FC = -3.46 | q.value = 0.0478 |
| 25 | EMPTY | RAB14 | IG51_AR1 | (PC9_IRTR) | (PC9_DMSO) | FC = 3.36 | q.value = 0.0309 |
| 26 | EMPTY | C2orf54 SNED1 | IG4_AR1 | (PC9_IRTR) | (PC9_DMSO) | FC = 3.10 | q.value = 0.0345 |
| 27 | EMPTY | AGBL5 EMILIN1 | IG26_AR3 | (PC9_IRTR) | (PC9_DMSO) | FC = 3.04 | q.value = 1.34e-05 |
| 28 | EMPTY | NACA | IG51_AR1 | (PC9_IRTR) | (PC9_DMSO) | FC = 3.00 | q.value = 7.68e-07 |
| 29 | EMPTY | AKR1C1 AKR1C2 | IG23_AR1 | (PC9_IRTR) | (PC9_DMSO) | FC = -2.92 | q.value = 0.0205 |
| 30 | EMPTY | SEC23B HARS2 | IG41_AR2 | (PC9_IRTR) | (PC9_DMSO) | FC = 2.83 | q.value = 0.0375 |
| 31 | EMPTY | RPL23 LASP1 | IG41_AR3 | (PC9_IRTR) | (PC9_DMSO) | FC = 2.64 | q.value = 0.0331 |
| 32 | EMPTY | AC114498.2 AC114498.2 | IG22_AR1 | (PC9_IRTR) | (PC9_DMSO) | FC = -2.11 | q.value = 1.9e-05 |
| 33 | EMPTY | FAM38A AC138028.1 | IG6_AR1 | (PC9_IRTR) | (PC9_DMSO) | FC = -2.11 | q.value = 0.017 |